Neural CDE¤
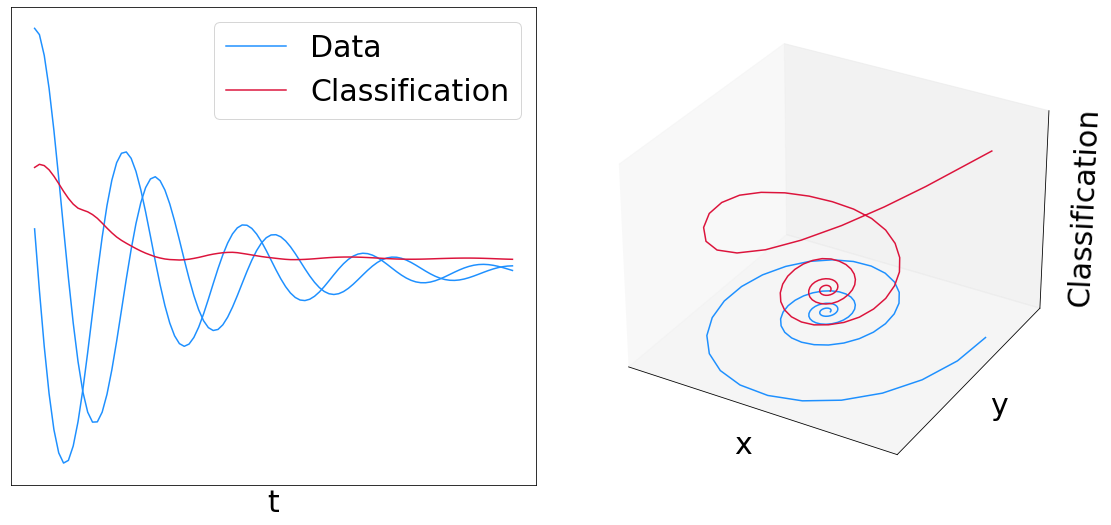
This example trains a Neural CDE (a "continuous time RNN") to distinguish clockwise from counter-clockwise spirals.
A neural CDE looks like
\(y(t) = y(0) + \int_0^t f_\theta(y(s)) \mathrm{d}x(s)\)
Where \(f_\theta\) is a neural network, and \(x\) is your data. The right hand side is a matrix-vector product between them. The integral is a Riemann--Stieltjes integral.
Info
Provided the path \(x\) is differentiable then the Riemann--Stieltjes integral can be converted into a normal integral:
\(y(t) = y(0) + \int_0^t f_\theta(y(s)) \frac{\mathrm{d}x}{\mathrm{d}s}(s) \mathrm{d}s\)
and in this case you can actually solve the CDE as an ODE. Indeed this is what we do below.
Typically the path \(x\) is constructed as a continuous interpolation of your input data. This is an approach that often makes a lot of sense when dealing with irregular data, densely sampled data etc. (i.e. the things that an RNN or Transformer might not work so well on.)
Reference:
@incollection{kidger2020neuralcde,
title={Neural Controlled Differential Equations for Irregular Time Series},
author={Kidger, Patrick and Morrill, James and Foster, James and Lyons, Terry},
booktitle={Advances in Neural Information Processing Systems},
publisher={Curran Associates, Inc.},
year={2020},
}
This example is available as a Jupyter notebook here.
import math
import time
import diffrax
import equinox as eqx # https://github.com/patrick-kidger/equinox
import jax
import jax.nn as jnn
import jax.numpy as jnp
import jax.random as jr
import jax.scipy as jsp
import matplotlib
import matplotlib.pyplot as plt
import optax # https://github.com/deepmind/optax
matplotlib.rcParams.update({"font.size": 30})
First let's define the vector field for the CDE.
class Func(eqx.Module):
mlp: eqx.nn.MLP
data_size: int
hidden_size: int
def __init__(self, data_size, hidden_size, width_size, depth, *, key, **kwargs):
super().__init__(**kwargs)
self.data_size = data_size
self.hidden_size = hidden_size
self.mlp = eqx.nn.MLP(
in_size=hidden_size,
out_size=hidden_size * data_size,
width_size=width_size,
depth=depth,
activation=jnn.softplus,
# Note the use of a tanh final activation function. This is important to
# stop the model blowing up. (Just like how GRUs and LSTMs constrain the
# rate of change of their hidden states.)
final_activation=jnn.tanh,
key=key,
)
def __call__(self, t, y, args):
return self.mlp(y).reshape(self.hidden_size, self.data_size)
Now wrap up the whole CDE solve into a model.
In this case we cap the neural CDE with a linear layer and sigmoid, to perform binary classification.
class NeuralCDE(eqx.Module):
initial: eqx.nn.MLP
func: Func
linear: eqx.nn.Linear
def __init__(self, data_size, hidden_size, width_size, depth, *, key, **kwargs):
super().__init__(**kwargs)
ikey, fkey, lkey = jr.split(key, 3)
self.initial = eqx.nn.MLP(data_size, hidden_size, width_size, depth, key=ikey)
self.func = Func(data_size, hidden_size, width_size, depth, key=fkey)
self.linear = eqx.nn.Linear(hidden_size, 1, key=lkey)
def __call__(self, ts, coeffs, evolving_out=False):
# Each sample of data consists of some timestamps `ts`, and some `coeffs`
# parameterising a control path. These are used to produce a continuous-time
# input path `control`.
control = diffrax.CubicInterpolation(ts, coeffs)
term = diffrax.ControlTerm(self.func, control).to_ode()
solver = diffrax.Tsit5()
dt0 = None
y0 = self.initial(control.evaluate(ts[0]))
if evolving_out:
saveat = diffrax.SaveAt(ts=ts)
else:
saveat = diffrax.SaveAt(t1=True)
solution = diffrax.diffeqsolve(
term,
solver,
ts[0],
ts[-1],
dt0,
y0,
stepsize_controller=diffrax.PIDController(rtol=1e-3, atol=1e-6),
saveat=saveat,
)
if evolving_out:
prediction = jax.vmap(lambda y: jnn.sigmoid(self.linear(y))[0])(solution.ys)
else:
(prediction,) = jnn.sigmoid(self.linear(solution.ys[-1]))
return prediction
Toy dataset of spirals.
We interpolate the samples with Hermite cubic splines with backward differences, which were introduced in https://arxiv.org/abs/2106.11028. (And produces better results than the natural cubic splines used in the original neural CDE paper.)
Time is a channel
Note the inclusion of time as a channel of the data! This is a subtle point that is often accidentally missed. If you include it then the model has enough information so that in theory it's actually a universal approximator. If you forget it then the model probably won't work very well...
If a CDE ever isn't training very well, make sure to ask yourself "did I include time as a channel?"
def get_data(dataset_size, add_noise, *, key):
theta_key, noise_key = jr.split(key, 2)
length = 100
theta = jr.uniform(theta_key, (dataset_size,), minval=0, maxval=2 * math.pi)
y0 = jnp.stack([jnp.cos(theta), jnp.sin(theta)], axis=-1)
ts = jnp.broadcast_to(jnp.linspace(0, 4 * math.pi, length), (dataset_size, length))
matrix = jnp.array([[-0.3, 2], [-2, -0.3]])
ys = jax.vmap(
lambda y0i, ti: jax.vmap(lambda tij: jsp.linalg.expm(tij * matrix) @ y0i)(ti)
)(y0, ts)
ys = jnp.concatenate([ts[:, :, None], ys], axis=-1) # time is a channel
ys = ys.at[: dataset_size // 2, :, 1].multiply(-1)
if add_noise:
ys = ys + jr.normal(noise_key, ys.shape) * 0.1
coeffs = jax.vmap(diffrax.backward_hermite_coefficients)(ts, ys)
labels = jnp.zeros((dataset_size,))
labels = labels.at[: dataset_size // 2].set(1.0)
_, _, data_size = ys.shape
return ts, coeffs, labels, data_size
def dataloader(arrays, batch_size, *, key):
dataset_size = arrays[0].shape[0]
assert all(array.shape[0] == dataset_size for array in arrays)
indices = jnp.arange(dataset_size)
while True:
perm = jr.permutation(key, indices)
(key,) = jr.split(key, 1)
start = 0
end = batch_size
while end < dataset_size:
batch_perm = perm[start:end]
yield tuple(array[batch_perm] for array in arrays)
start = end
end = start + batch_size
The main entry point. Try running main() to train the neural CDE.
def main(
dataset_size=256,
add_noise=False,
batch_size=32,
lr=1e-2,
steps=20,
hidden_size=8,
width_size=128,
depth=1,
seed=5678,
):
key = jr.PRNGKey(seed)
train_data_key, test_data_key, model_key, loader_key = jr.split(key, 4)
ts, coeffs, labels, data_size = get_data(
dataset_size, add_noise, key=train_data_key
)
model = NeuralCDE(data_size, hidden_size, width_size, depth, key=model_key)
# Training loop like normal.
@eqx.filter_jit
def loss(model, ti, label_i, coeff_i):
pred = jax.vmap(model)(ti, coeff_i)
# Binary cross-entropy
bxe = label_i * jnp.log(pred) + (1 - label_i) * jnp.log(1 - pred)
bxe = -jnp.mean(bxe)
acc = jnp.mean((pred > 0.5) == (label_i == 1))
return bxe, acc
grad_loss = eqx.filter_value_and_grad(loss, has_aux=True)
@eqx.filter_jit
def make_step(model, data_i, opt_state):
ti, label_i, *coeff_i = data_i
(bxe, acc), grads = grad_loss(model, ti, label_i, coeff_i)
updates, opt_state = optim.update(grads, opt_state)
model = eqx.apply_updates(model, updates)
return bxe, acc, model, opt_state
optim = optax.adam(lr)
opt_state = optim.init(eqx.filter(model, eqx.is_inexact_array))
for step, data_i in zip(
range(steps), dataloader((ts, labels) + coeffs, batch_size, key=loader_key)
):
start = time.time()
bxe, acc, model, opt_state = make_step(model, data_i, opt_state)
end = time.time()
print(
f"Step: {step}, Loss: {bxe}, Accuracy: {acc}, Computation time: "
f"{end - start}"
)
ts, coeffs, labels, _ = get_data(dataset_size, add_noise, key=test_data_key)
bxe, acc = loss(model, ts, labels, coeffs)
print(f"Test loss: {bxe}, Test Accuracy: {acc}")
# Plot results
sample_ts = ts[-1]
sample_coeffs = tuple(c[-1] for c in coeffs)
pred = model(sample_ts, sample_coeffs, evolving_out=True)
interp = diffrax.CubicInterpolation(sample_ts, sample_coeffs)
values = jax.vmap(interp.evaluate)(sample_ts)
fig = plt.figure(figsize=(16, 8))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2, projection="3d")
ax1.plot(sample_ts, values[:, 1], c="dodgerblue")
ax1.plot(sample_ts, values[:, 2], c="dodgerblue", label="Data")
ax1.plot(sample_ts, pred, c="crimson", label="Classification")
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_xlabel("t")
ax1.legend()
ax2.plot(values[:, 1], values[:, 2], c="dodgerblue", label="Data")
ax2.plot(values[:, 1], values[:, 2], pred, c="crimson", label="Classification")
ax2.set_xticks([])
ax2.set_yticks([])
ax2.set_zticks([])
ax2.set_xlabel("x")
ax2.set_ylabel("y")
ax2.set_zlabel("Classification")
plt.tight_layout()
plt.savefig("neural_cde.png")
plt.show()
main()
Step: 0, Loss: 2.5234897136688232, Accuracy: 0.5, Computation time: 27.177752256393433
Step: 1, Loss: 4.682699203491211, Accuracy: 0.5, Computation time: 0.5112535953521729
Step: 2, Loss: 1.9817578792572021, Accuracy: 0.46875, Computation time: 0.4303276538848877
Step: 3, Loss: 0.909335732460022, Accuracy: 0.375, Computation time: 0.42275118827819824
Step: 4, Loss: 0.5238552093505859, Accuracy: 0.96875, Computation time: 0.3412055969238281
Step: 5, Loss: 0.5987676382064819, Accuracy: 0.5625, Computation time: 0.4041574001312256
Step: 6, Loss: 0.5615957975387573, Accuracy: 0.5625, Computation time: 0.3387322425842285
Step: 7, Loss: 0.5031553506851196, Accuracy: 0.625, Computation time: 0.4076976776123047
Step: 8, Loss: 0.3657313883304596, Accuracy: 0.84375, Computation time: 0.35105156898498535
Step: 9, Loss: 0.34929466247558594, Accuracy: 0.9375, Computation time: 0.42032384872436523
Step: 10, Loss: 0.2539682686328888, Accuracy: 1.0, Computation time: 0.3486146926879883
Step: 11, Loss: 0.2294737994670868, Accuracy: 1.0, Computation time: 0.3518819808959961
Step: 12, Loss: 0.2001168429851532, Accuracy: 1.0, Computation time: 0.4245719909667969
Step: 13, Loss: 0.18462520837783813, Accuracy: 1.0, Computation time: 0.4051353931427002
Step: 14, Loss: 0.19849714636802673, Accuracy: 1.0, Computation time: 0.4198932647705078
Step: 15, Loss: 0.21601906418800354, Accuracy: 1.0, Computation time: 0.344160795211792
Step: 16, Loss: 0.1362144500017166, Accuracy: 1.0, Computation time: 0.42815589904785156
Step: 17, Loss: 0.12172335386276245, Accuracy: 1.0, Computation time: 0.3531978130340576
Step: 18, Loss: 0.13752871751785278, Accuracy: 1.0, Computation time: 0.4119846820831299
Step: 19, Loss: 0.10557006299495697, Accuracy: 1.0, Computation time: 0.33621764183044434
Test loss: 0.1057077944278717, Test Accuracy: 1.0