Optimising a parameterised ODE¤
Here's an example in which we optimise the parameters of an ODE. This means we have a differential equation solve inside our nonlinear solve!
First, import everything. Regarding the libraries we're using: Diffrax for differential equation solving, jaxtyping for type annotations, and Equinox for some "smarter" JAX operations. (Equinox is used as the foundation of all of Optimistix/Diffrax/etc.)
import diffrax as dfx # https://github.com/patrick-kidger/diffrax
import equinox as eqx # https://github.com/patrick-kidger/equinox
import jax.numpy as jnp
import matplotlib.pyplot as plt
import optimistix as optx
from jaxtyping import Array, Float # https://github.com/google/jaxtyping
The problem we'll try and tackle is a relatively simple one: fitting the Lotka--Volterra equations.
def vector_field(
t, y: Float[Array, "2"], parameters: Float[Array, "4"]
) -> Float[Array, "2"]:
prey, predator = y
α, β, γ, δ = parameters
d_prey = α * prey - β * prey * predator
d_predator = -γ * predator + δ * prey * predator
d_y = jnp.stack([d_prey, d_predator])
return d_y
def solve(
parameters: Float[Array, "4"], y0: Float[Array, "2"], saveat: dfx.SaveAt
) -> Float[Array, "ts"]:
"""Solve a single ODE."""
term = dfx.ODETerm(vector_field)
solver = dfx.Tsit5()
t0 = saveat.subs.ts[0]
t1 = saveat.subs.ts[-1]
dt0 = 0.1
sol = dfx.diffeqsolve(
term,
solver,
t0,
t1,
dt0,
y0,
args=parameters,
saveat=saveat,
# support forward-mode autodiff, which is used by Levenberg--Marquardt
adjoint=dfx.DirectAdjoint(),
)
return sol.ys
Let's quickly simulate some training data. The fact that this is also coming from the Lotka--Volterra equations means that we're hoping to get a perfect fit with the parameters. In practice, on real-world data, that's unlikely to be true!
def get_data() -> tuple[Float[Array, "3 2"], Float[Array, "3 50"]]:
"""Simulate some training data."""
# We consider three possible initial conditions.
y0_a = jnp.array([9.0, 9.0])
y0_b = jnp.array([10.0, 10.0])
y0_c = jnp.array([11.0, 11.0])
y0s = jnp.stack([y0_a, y0_b, y0_c])
true_parameters = jnp.array([0.1, 0.02, 0.4, 0.02])
saveat = dfx.SaveAt(ts=jnp.linspace(0, 30, 20))
batch_solve = eqx.filter_jit(eqx.filter_vmap(solve, in_axes=(None, 0, None)))
values = batch_solve(true_parameters, y0s, saveat)
return y0s, values
And now let's run our solver.
Some solvers support a verbose option, to get a printout of how the optimisation proceeds. We turn that on in this case.
def residuals(parameters, y0s__values):
y0s, values = y0s__values
saveat = dfx.SaveAt(ts=jnp.linspace(0, 30, 20))
batch_solve = eqx.filter_vmap(solve, in_axes=(None, 0, None))
pred_values = batch_solve(parameters, y0s, saveat)
return values - pred_values
(y0s, values) = get_data()
solver = optx.LevenbergMarquardt(rtol=1e-8, atol=1e-8, verbose=True)
init_parameters = jnp.zeros(4)
sol = optx.least_squares(residuals, solver, init_parameters, args=(y0s, values))
Step: 0 , Accepted steps: 0 , Steps since acceptance: 0 , Loss on this step: 8342.5693359375 , Loss on the last accepted step: 0.0 , Step size: 1.0
Step: 1 , Accepted steps: 1 , Steps since acceptance: 0 , Loss on this step: 8628328.0 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.25
Step: 2 , Accepted steps: 1 , Steps since acceptance: 1 , Loss on this step: 8608194.0 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.0625
Step: 3 , Accepted steps: 1 , Steps since acceptance: 2 , Loss on this step: 8528131.0 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.015625
Step: 4 , Accepted steps: 1 , Steps since acceptance: 3 , Loss on this step: 8217127.5 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.00390625
Step: 5 , Accepted steps: 1 , Steps since acceptance: 4 , Loss on this step: 7105586.5 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.0009765625
Step: 6 , Accepted steps: 1 , Steps since acceptance: 5 , Loss on this step: 4168893.0 , Loss on the last accepted step: 8342.5693359375 , Step size: 0.000244140625
Step: 7 , Accepted steps: 1 , Steps since acceptance: 6 , Loss on this step: 859413.5625 , Loss on the last accepted step: 8342.5693359375 , Step size: 6.103515625e-05
Step: 8 , Accepted steps: 1 , Steps since acceptance: 7 , Loss on this step: 51456.02734375 , Loss on the last accepted step: 8342.5693359375 , Step size: 1.52587890625e-05
Step: 9 , Accepted steps: 1 , Steps since acceptance: 8 , Loss on this step: 4537.26025390625 , Loss on the last accepted step: 8342.5693359375 , Step size: 1.52587890625e-05
Step: 10 , Accepted steps: 2 , Steps since acceptance: 0 , Loss on this step: 1933.3726806640625 , Loss on the last accepted step: 4537.26025390625 , Step size: 1.52587890625e-05
Step: 11 , Accepted steps: 3 , Steps since acceptance: 0 , Loss on this step: 1527.572021484375 , Loss on the last accepted step: 1933.3726806640625 , Step size: 1.52587890625e-05
Step: 12 , Accepted steps: 4 , Steps since acceptance: 0 , Loss on this step: 1475.835205078125 , Loss on the last accepted step: 1527.572021484375 , Step size: 5.340576171875e-05
Step: 13 , Accepted steps: 5 , Steps since acceptance: 0 , Loss on this step: 1446.6658935546875 , Loss on the last accepted step: 1475.835205078125 , Step size: 0.000186920166015625
Step: 14 , Accepted steps: 6 , Steps since acceptance: 0 , Loss on this step: 1423.8236083984375 , Loss on the last accepted step: 1446.6658935546875 , Step size: 0.0006542205810546875
Step: 15 , Accepted steps: 7 , Steps since acceptance: 0 , Loss on this step: 1345.58154296875 , Loss on the last accepted step: 1423.8236083984375 , Step size: 0.0022897720336914062
Step: 16 , Accepted steps: 8 , Steps since acceptance: 0 , Loss on this step: 12441.5048828125 , Loss on the last accepted step: 1345.58154296875 , Step size: 0.0005724430084228516
Step: 17 , Accepted steps: 8 , Steps since acceptance: 1 , Loss on this step: 9109.619140625 , Loss on the last accepted step: 1345.58154296875 , Step size: 0.0001431107521057129
Step: 18 , Accepted steps: 8 , Steps since acceptance: 2 , Loss on this step: 5108.0771484375 , Loss on the last accepted step: 1345.58154296875 , Step size: 3.577768802642822e-05
Step: 19 , Accepted steps: 8 , Steps since acceptance: 3 , Loss on this step: 4064.636962890625 , Loss on the last accepted step: 1345.58154296875 , Step size: 8.944422006607056e-06
Step: 20 , Accepted steps: 8 , Steps since acceptance: 4 , Loss on this step: 3757.67236328125 , Loss on the last accepted step: 1345.58154296875 , Step size: 2.236105501651764e-06
Step: 21 , Accepted steps: 8 , Steps since acceptance: 5 , Loss on this step: 3031.880859375 , Loss on the last accepted step: 1345.58154296875 , Step size: 5.59026375412941e-07
Step: 22 , Accepted steps: 8 , Steps since acceptance: 6 , Loss on this step: 1061.90771484375 , Loss on the last accepted step: 1345.58154296875 , Step size: 1.9565923139452934e-06
Step: 23 , Accepted steps: 9 , Steps since acceptance: 0 , Loss on this step: 624.8424682617188 , Loss on the last accepted step: 1061.90771484375 , Step size: 1.9565923139452934e-06
Step: 24 , Accepted steps: 10 , Steps since acceptance: 0 , Loss on this step: 361.8550720214844 , Loss on the last accepted step: 624.8424682617188 , Step size: 6.848073098808527e-06
Step: 25 , Accepted steps: 11 , Steps since acceptance: 0 , Loss on this step: 122.06681060791016 , Loss on the last accepted step: 361.8550720214844 , Step size: 2.3968255845829844e-05
Step: 26 , Accepted steps: 12 , Steps since acceptance: 0 , Loss on this step: 38.64931106567383 , Loss on the last accepted step: 122.06681060791016 , Step size: 2.3968255845829844e-05
Step: 27 , Accepted steps: 13 , Steps since acceptance: 0 , Loss on this step: 21.20478630065918 , Loss on the last accepted step: 38.64931106567383 , Step size: 2.3968255845829844e-05
Step: 28 , Accepted steps: 14 , Steps since acceptance: 0 , Loss on this step: 15.529594421386719 , Loss on the last accepted step: 21.20478630065918 , Step size: 2.3968255845829844e-05
Step: 29 , Accepted steps: 15 , Steps since acceptance: 0 , Loss on this step: 11.561433792114258 , Loss on the last accepted step: 15.529594421386719 , Step size: 2.3968255845829844e-05
Step: 30 , Accepted steps: 16 , Steps since acceptance: 0 , Loss on this step: 8.713208198547363 , Loss on the last accepted step: 11.561433792114258 , Step size: 2.3968255845829844e-05
Step: 31 , Accepted steps: 17 , Steps since acceptance: 0 , Loss on this step: 6.632508754730225 , Loss on the last accepted step: 8.713208198547363 , Step size: 8.388889546040446e-05
Step: 32 , Accepted steps: 18 , Steps since acceptance: 0 , Loss on this step: 3.073559284210205 , Loss on the last accepted step: 6.632508754730225 , Step size: 8.388889546040446e-05
Step: 33 , Accepted steps: 19 , Steps since acceptance: 0 , Loss on this step: 1.434643030166626 , Loss on the last accepted step: 3.073559284210205 , Step size: 8.388889546040446e-05
Step: 34 , Accepted steps: 20 , Steps since acceptance: 0 , Loss on this step: 0.6955414414405823 , Loss on the last accepted step: 1.434643030166626 , Step size: 0.0002936111413873732
Step: 35 , Accepted steps: 21 , Steps since acceptance: 0 , Loss on this step: 0.12273844331502914 , Loss on the last accepted step: 0.6955414414405823 , Step size: 0.0002936111413873732
Step: 36 , Accepted steps: 22 , Steps since acceptance: 0 , Loss on this step: 0.02028850093483925 , Loss on the last accepted step: 0.12273844331502914 , Step size: 0.0010276390239596367
Step: 37 , Accepted steps: 23 , Steps since acceptance: 0 , Loss on this step: 0.0006190080312080681 , Loss on the last accepted step: 0.02028850093483925 , Step size: 0.0035967365838587284
Step: 38 , Accepted steps: 24 , Steps since acceptance: 0 , Loss on this step: 1.9843394056806574e-06 , Loss on the last accepted step: 0.0006190080312080681 , Step size: 0.012588578276336193
Step: 39 , Accepted steps: 25 , Steps since acceptance: 0 , Loss on this step: 6.252315998267477e-09 , Loss on the last accepted step: 1.9843394056806574e-06 , Step size: 0.04406002536416054
Step: 40 , Accepted steps: 26 , Steps since acceptance: 0 , Loss on this step: 1.3584919855702537e-09 , Loss on the last accepted step: 6.252315998267477e-09 , Step size: 0.15421009063720703
Step: 41 , Accepted steps: 27 , Steps since acceptance: 0 , Loss on this step: 1.2424514750364324e-09 , Loss on the last accepted step: 1.3584919855702537e-09 , Step size: 0.15421009063720703
Step: 42 , Accepted steps: 28 , Steps since acceptance: 0 , Loss on this step: 2.684974020894515e-09 , Loss on the last accepted step: 1.2424514750364324e-09 , Step size: 0.03855252265930176
Step: 43 , Accepted steps: 28 , Steps since acceptance: 1 , Loss on this step: 2.684974020894515e-09 , Loss on the last accepted step: 1.2424514750364324e-09 , Step size: 0.00963813066482544
Step: 44 , Accepted steps: 28 , Steps since acceptance: 2 , Loss on this step: 2.684974020894515e-09 , Loss on the last accepted step: 1.2424514750364324e-09 , Step size: 0.00240953266620636
Step: 45 , Accepted steps: 28 , Steps since acceptance: 3 , Loss on this step: 2.684974020894515e-09 , Loss on the last accepted step: 1.2424514750364324e-09 , Step size: 0.00060238316655159
Step: 46 , Accepted steps: 28 , Steps since acceptance: 4 , Loss on this step: 8.882565794010588e-10 , Loss on the last accepted step: 1.2424514750364324e-09 , Step size: 0.00060238316655159
Step: 47 , Accepted steps: 29 , Steps since acceptance: 0 , Loss on this step: 1.7594743439985905e-09 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 0.0001505957916378975
Step: 48 , Accepted steps: 29 , Steps since acceptance: 1 , Loss on this step: 2.120401632055291e-09 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 3.764894790947437e-05
Step: 49 , Accepted steps: 29 , Steps since acceptance: 2 , Loss on this step: 2.0302213243894585e-09 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 9.412236977368593e-06
Step: 50 , Accepted steps: 29 , Steps since acceptance: 3 , Loss on this step: 9.237606235501516e-10 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 2.3530592443421483e-06
Step: 51 , Accepted steps: 29 , Steps since acceptance: 4 , Loss on this step: 9.347882468091484e-10 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 5.882648110855371e-07
Step: 52 , Accepted steps: 29 , Steps since acceptance: 5 , Loss on this step: 9.347882468091484e-10 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 1.4706620277138427e-07
Step: 53 , Accepted steps: 29 , Steps since acceptance: 6 , Loss on this step: 9.347882468091484e-10 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 3.676655069284607e-08
Step: 54 , Accepted steps: 29 , Steps since acceptance: 7 , Loss on this step: 8.882565794010588e-10 , Loss on the last accepted step: 8.882565794010588e-10 , Step size: 3.676655069284607e-08
Alright, we're getting told that the loss is about zero. Let's check that:
optx.max_norm(residuals(sol.value, (y0s, values)))
Array(1.9073486e-05, dtype=float32)
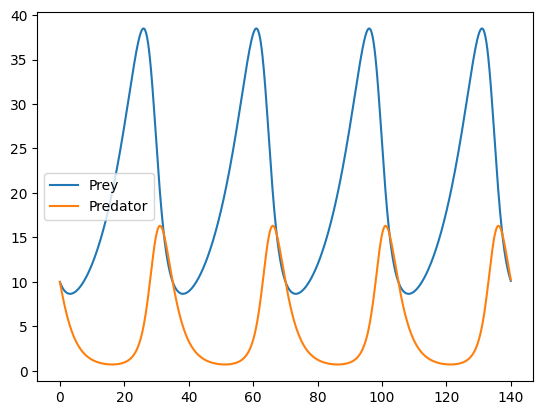
That looks pretty good! Let's see what the solution looks like:
ts = jnp.linspace(0, 140, 1000)
ys = solve(sol.value, jnp.array([10.0, 10.0]), dfx.SaveAt(ts=ts))
plt.plot(ts, ys[:, 0], label="Prey")
plt.plot(ts, ys[:, 1], label="Predator")
plt.legend()
plt.show()
Tip
Here are two tips for fitting ODEs like this.
First, we started with an initial guess of all of our parameters being zero. This corresponds to the vector field of the ODE being zero, which means that the initial solution of the ODE is constant. In contrast if we'd set some other values for the initial parameters, then our ODE might evolve to produce trajectories that are very far away from our data.
Second: we fit to quite a short time series (20 points over the interval [0, 30]; note that we produced our final plot over a much longer time interval). It's quite common for optimisation to get stuck in a bad local minima if you try to fit to a long time series straight away. It's much better to gradually lengthen the time series you try to fit.